How to make your machine learning model available as an API with the plumber package

The plumber package for R makes it easy to expose existing R code as a webservice via an API (https://www.rplumber.io/, Trestle Technology, LLC 2017).
You take an existing R script and make it accessible with plumber by simply adding a few lines of comments. If you have worked with Roxygen before, e.g. when building a package, you will already be familiar with the core concepts. If not, here are the most important things to know:
- you define the output or endpoint
- you can add additional annotation to customize your input, output and other functionalities of your API
- you can define every input parameter that will go into your function
- every such annotation will begin with either
#'or#*
With this setup, we can take a trained machine learning model and make it available as an API. With this API, other programs can access it and use it to make predictions.
What are APIs and webservices?
With plumber, we can build so called HTTP APIs. HTTP stands for Hypertext Transfer Protocol and is used to transmit information on the web; API stands for Application Programming Interface and governs the connection between some software and underlying applications. Software can then communicate via HTTP APIs. This way, our R script can be called from other software, even if the other program is not written in R and we have built a tool for machine-to-machine communication, i.e. a webservice.
How to convert your R script into an API with plumber
Training and saving a model
Let’s say we have trained a machine learning model as in this post about LIME. I loaded a data set on chronic kidney disease, did some preprocessing (converting categorical features into dummy variables, scaling and centering), split it into training and test data and trained a Random Forest model with caret. We can use this trained model to make predictions for one test case with the following code:
library(tidyverse)
# load test and train data
load("../../data/test_data.RData")
load("../../data/train_data.RData")
# load model
load("../../data/model_rf.RData")
# take first test case for prediction
input_data <- test_data[1, ] %>%
select(-class)
# predict test case using model
pred <- predict(model_rf, input_data)
cat("----------------\nTest case predicted to be", as.character(pred), "\n----------------")## ----------------
## Test case predicted to be ckd
## ----------------The input
For our API to work, we need to define the input, in our case the features of the test data. When we look at the model object, we see that it expects the following parameters:
var_names <- model_rf$finalModel$xNames
var_names## [1] "age" "bp" "sg_1.005" "sg_1.010"
## [5] "sg_1.015" "sg_1.020" "sg_1.025" "al_0"
## [9] "al_1" "al_2" "al_3" "al_4"
## [13] "al_5" "su_0" "su_1" "su_2"
## [17] "su_3" "su_4" "su_5" "rbc_normal"
## [21] "rbc_abnormal" "pc_normal" "pc_abnormal" "pcc_present"
## [25] "pcc_notpresent" "ba_present" "ba_notpresent" "bgr"
## [29] "bu" "sc" "sod" "pot"
## [33] "hemo" "pcv" "wbcc" "rbcc"
## [37] "htn_yes" "htn_no" "dm_yes" "dm_no"
## [41] "cad_yes" "cad_no" "appet_good" "appet_poor"
## [45] "pe_yes" "pe_no" "ane_yes" "ane_no"Good practice is to write the input parameter definition into you API Swagger UI, but the code would work without these annotations. We define the parameters by annotating them with name and description in our R-script using @parameter. For this purpose, I want to know the type and min/max values for each of my variables in the training data. Because categorical data has been converted to dummy variables and then scaled and centered, these values will all be numeric and between 0 and 1 in this example. If I would build this script for a real case, I’d use the raw data as input and add a preprocessing function to my script, though!
# show parameter definition for the first three features
for (i in 1:3) {
# if you wanted to see it for all features, use
#for (i in 1:length(var_names)) {
var <- var_names[i]
train_data_subs <- train_data[, which(colnames(train_data) == var)]
type <- class(train_data_subs)
if (type == "numeric") {
min <- min(train_data_subs)
max <- max(train_data_subs)
}
cat("Variable:", var, "is of type:", type, "\n",
"Min value in training data =", min, "\n",
"Max value in training data =", max, "\n----------\n")
}## Variable: age is of type: numeric
## Min value in training data = 0
## Max value in training data = 0.9777778
## ----------
## Variable: bp is of type: numeric
## Min value in training data = 0
## Max value in training data = 0.7222222
## ----------
## Variable: sg_1.005 is of type: numeric
## Min value in training data = 0
## Max value in training data = 1
## ----------Unless otherwise instructed, all parameters passed into plumber endpoints from query strings or dynamic paths will be character strings. https://www.rplumber.io/docs/routing-and-input.html#typed-dynamic-routes
This means that we need to convert numeric values before we process them further. Or we can define the parameter type explicitly, e.g. by writing variable_1:numeric if we want to specifiy that variable_1 is supposed to be numeric.
To make sure that the model will perform as expected, it is also advisable to add a few validation functions. Here, I will validate
- whether every parameter is numeric/integer by checking for NAs (which would have resulted from
as.numeric()/as.integer()applied to data of character type) - whether every parameter is between 0 and 1
In order for plumber to work with our input, it needs to be part of the HTTP request, which can then be routed to our R function. The plumber documentation describes how to use query strings as inputs. But in our case, manually writing query strings is not practical because we have so many parameters. Of course, there are programs that let us generate query strings but the easiest way to format the input from a line of data I found is to use JSON.
The toJSON() function from the rjson package converts our input line to JSON format:
library(rjson)
test_case_json <- toJSON(input_data)
cat(test_case_json)## {"age":0.511111111111111,"bp":0.111111111111111,"sg_1.005":1,"sg_1.010":0,"sg_1.015":0,"sg_1.020":0,"sg_1.025":0,"al_0":0,"al_1":0,"al_2":0,"al_3":0,"al_4":1,"al_5":0,"su_0":1,"su_1":0,"su_2":0,"su_3":0,"su_4":0,"su_5":0,"rbc_normal":1,"rbc_abnormal":0,"pc_normal":0,"pc_abnormal":1,"pcc_present":1,"pcc_notpresent":0,"ba_present":0,"ba_notpresent":1,"bgr":0.193877551020408,"bu":0.139386189258312,"sc":0.0447368421052632,"sod":0.653374233128834,"pot":0,"hemo":0.455056179775281,"pcv":0.425925925925926,"wbcc":0.170454545454545,"rbcc":0.225,"htn_yes":1,"htn_no":0,"dm_yes":0,"dm_no":1,"cad_yes":0,"cad_no":1,"appet_good":0,"appet_poor":1,"pe_yes":1,"pe_no":0,"ane_yes":1,"ane_no":0}Defining the endpoint and output
In order to convert this very simple script into an API, we need to define the endpoint(s). Endpoints will return an output, in our case it will return the output of the predict() function pasted into a line of text (e.g. “Test case predicted to be ckd”). Here, we want to have the predictions returned, so we annotate the entire function with @get. This endpoint in the API will get a custom name, so that we can call it later; here we call it predict and therefore write #' @get /predict.
According to the design of the HTTP specification, GET (along with HEAD) requests are used only to read data and not change it. Therefore, when used this way, they are considered safe. That is, they can be called without risk of data modification or corruption — calling it once has the same effect as calling it 10 times, or none at all. Additionally, GET (and HEAD) is idempotent, which means that making multiple identical requests ends up having the same result as a single request. http://www.restapitutorial.com/lessons/httpmethods.html
In this case, we could also consider using @post to avoid caching issues, but for this example I’ll leave it as @get.
The POST verb is most-often utilized to create new resources. In particular, it’s used to create subordinate resources. That is, subordinate to some other (e.g. parent) resource. In other words, when creating a new resource, POST to the parent and the service takes care of associating the new resource with the parent, assigning an ID (new resource URI), etc. On successful creation, return HTTP status 201, returning a Location header with a link to the newly-created resource with the 201 HTTP status. POST is neither safe nor idempotent. It is therefore recommended for non-idempotent resource requests. Making two identical POST requests will most-likely result in two resources containing the same information. http://www.restapitutorial.com/lessons/httpmethods.html
We can also customize the output. Keep in mind though, that the output should be “serialized”. By default, the output will be in JSON format. Here, I want to have a text output, so I’ll specify @html without html formatting specifications, although I could add them if I wanted to display the text on a website. If we were to store the data in a database, however, this would not be a good idea. In that case, it would be better to output the result as a JSON object.
Logging with filters
It is also useful to provide some sort of logging for your API. Here, I am using the simple example from the plumber documentation that uses filters and output the logs to the console or your API server logs. You could also write your logging output to a file. In production, it would be better to use a real logging setup that stores information about each request, e.g. the time stamp, whether any errors or warnings occurred, etc. The forward() part of the logging function passes control on to the next handler in the pipeline, here our predict function.
Running the plumber script
We need to save the entire script with annotations as an .R file as seen below. The regular comments # describe what each section does.
# script name:
# plumber.R
# set API title and description to show up in http://localhost:8000/__swagger__/
#' @apiTitle Run predictions for Chronic Kidney Disease with Random Forest Model
#' @apiDescription This API takes as patient data on Chronic Kidney Disease and returns a prediction whether the lab values
#' indicate Chronic Kidney Disease (ckd) or not (notckd).
#' For details on how the model is built, see https://shirinsplayground.netlify.com/2017/12/lime_sketchnotes/
#' For further explanations of this plumber function, see https://shirinsplayground.netlify.com/2018/01/plumber/
# load model
# this path would have to be adapted if you would deploy this
load("/Users/shiringlander/Documents/Github/shirinsplayground/data/model_rf.RData")
#' Log system time, request method and HTTP user agent of the incoming request
#' @filter logger
function(req){
cat("System time:", as.character(Sys.time()), "\n",
"Request method:", req$REQUEST_METHOD, req$PATH_INFO, "\n",
"HTTP user agent:", req$HTTP_USER_AGENT, "@", req$REMOTE_ADDR, "\n")
plumber::forward()
}
# core function follows below:
# define parameters with type and description
# name endpoint
# return output as html/text
# specify 200 (okay) return
#' predict Chronic Kidney Disease of test case with Random Forest model
#' @param age:numeric The age of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param bp:numeric The blood pressure of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param sg_1.005:int The urinary specific gravity of the patient, integer (1: sg = 1.005, otherwise 0)
#' @param sg_1.010:int The urinary specific gravity of the patient, integer (1: sg = 1.010, otherwise 0)
#' @param sg_1.015:int The urinary specific gravity of the patient, integer (1: sg = 1.015, otherwise 0)
#' @param sg_1.020:int The urinary specific gravity of the patient, integer (1: sg = 1.020, otherwise 0)
#' @param sg_1.025:int The urinary specific gravity of the patient, integer (1: sg = 1.025, otherwise 0)
#' @param al_0:int The urine albumin level of the patient, integer (1: al = 0, otherwise 0)
#' @param al_1:int The urine albumin level of the patient, integer (1: al = 1, otherwise 0)
#' @param al_2:int The urine albumin level of the patient, integer (1: al = 2, otherwise 0)
#' @param al_3:int The urine albumin level of the patient, integer (1: al = 3, otherwise 0)
#' @param al_4:int The urine albumin level of the patient, integer (1: al = 4, otherwise 0)
#' @param al_5:int The urine albumin level of the patient, integer (1: al = 5, otherwise 0)
#' @param su_0:int The sugar level of the patient, integer (1: su = 0, otherwise 0)
#' @param su_1:int The sugar level of the patient, integer (1: su = 1, otherwise 0)
#' @param su_2:int The sugar level of the patient, integer (1: su = 2, otherwise 0)
#' @param su_3:int The sugar level of the patient, integer (1: su = 3, otherwise 0)
#' @param su_4:int The sugar level of the patient, integer (1: su = 4, otherwise 0)
#' @param su_5:int The sugar level of the patient, integer (1: su = 5, otherwise 0)
#' @param rbc_normal:int The red blood cell count of the patient, integer (1: rbc = normal, otherwise 0)
#' @param rbc_abnormal:int The red blood cell count of the patient, integer (1: rbc = abnormal, otherwise 0)
#' @param pc_normal:int The pus cell level of the patient, integer (1: pc = normal, otherwise 0)
#' @param pc_abnormal:int The pus cell level of the patient, integer (1: pc = abnormal, otherwise 0)
#' @param pcc_present:int The puc cell clumps status of the patient, integer (1: pcc = present, otherwise 0)
#' @param pcc_notpresent:int The puc cell clumps status of the patient, integer (1: pcc = notpresent, otherwise 0)
#' @param ba_present:int The bacteria status of the patient, integer (1: ba = present, otherwise 0)
#' @param ba_notpresent:int The bacteria status of the patient, integer (1: ba = notpresent, otherwise 0)
#' @param bgr:numeric The blood glucose random level of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param bu:numeric The blood urea level of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param sc:numeric The serum creatinine level of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param sod:numeric The sodium level of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param pot:numeric The potassium level of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param hemo:numeric The hemoglobin level of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param pcv:numeric The packed cell volume of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param wbcc:numeric The white blood cell count of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param rbcc:numeric The red blood cell count of the patient, numeric (scaled and centered to be btw 0 and 1)
#' @param htn_yes:int The hypertension status of the patient, integer (1: htn = yes, otherwise 0)
#' @param htn_no:int The hypertension status of the patient, integer (1: htn = no, otherwise 0)
#' @param dm_yes:int The diabetes mellitus status of the patient, integer (1: dm = yes, otherwise 0)
#' @param dm_no:int The diabetes mellitus status of the patient, integer (1: dm = no, otherwise 0)
#' @param cad_yes:int The coronary artery disease status of the patient, integer (1: cad = yes, otherwise 0)
#' @param cad_no:int The coronary artery disease status of the patient, integer (1: cad = no, otherwise 0)
#' @param appet_good:int The appetite of the patient, integer (1: appet = good, otherwise 0)
#' @param appet_poor:int The appetite of the patient, integer (1: appet = poor, otherwise 0)
#' @param pe_yes:int The pedal edema status of the patient, integer (1: pe = yes, otherwise 0)
#' @param pe_no:int The pedal edema status of the patient, integer (1: pe = no, otherwise 0)
#' @param ane_yes:int The anemia status of the patient, integer (1: ane = yes, otherwise 0)
#' @param ane_no:int The anemia status of the patient, integer (1: ane = no, otherwise 0)
#' @get /predict
#' @html
#' @response 200 Returns the class (ckd or notckd) prediction from the Random Forest model; ckd = Chronic Kidney Disease
calculate_prediction <- function(age, bp, sg_1.005, sg_1.010, sg_1.015, sg_1.020, sg_1.025, al_0, al_1, al_2,
al_3, al_4, al_5, su_0, su_1, su_2, su_3, su_4, su_5, rbc_normal, rbc_abnormal, pc_normal, pc_abnormal,
pcc_present, pcc_notpresent, ba_present, ba_notpresent, bgr, bu, sc, sod, pot, hemo, pcv,
wbcc, rbcc, htn_yes, htn_no, dm_yes, dm_no, cad_yes, cad_no, appet_good, appet_poor, pe_yes, pe_no,
ane_yes, ane_no) {
# make data frame from numeric parameters
input_data_num <<- data.frame(age, bp, bgr, bu, sc, sod, pot, hemo, pcv, wbcc, rbcc,
stringsAsFactors = FALSE)
# and make sure they really are numeric
input_data_num <<- as.data.frame(t(sapply(input_data_num, as.numeric)))
# make data frame from (binary) integer parameters
input_data_int <<- data.frame(sg_1.005, sg_1.010, sg_1.015, sg_1.020, sg_1.025, al_0, al_1, al_2,
al_3, al_4, al_5, su_0, su_1, su_2, su_3, su_4, su_5, rbc_normal, rbc_abnormal, pc_normal, pc_abnormal,
pcc_present, pcc_notpresent, ba_present, ba_notpresent, htn_yes, htn_no, dm_yes, dm_no,
cad_yes, cad_no, appet_good, appet_poor, pe_yes, pe_no, ane_yes, ane_no,
stringsAsFactors = FALSE)
# and make sure they really are numeric
input_data_int <<- as.data.frame(t(sapply(input_data_int, as.integer)))
# combine into one data frame
input_data <<- as.data.frame(cbind(input_data_num, input_data_int))
# validation for parameter
if (any(is.na(input_data))) {
res$status <- 400
res$body <- "Parameters have to be numeric or integers"
}
if (any(input_data < 0) || any(input_data > 1)) {
res$status <- 400
res$body <- "Parameters have to be between 0 and 1"
}
# predict and return result
pred_rf <<- predict(model_rf, input_data)
paste("----------------\nTest case predicted to be", as.character(pred_rf), "\n----------------\n")
}
Note that I am using the “double-assignment” operator <<- in my function, because I want to make sure that objects are overwritten at the top level (i.e. globally). This would have been relevant had I set a global parameter, but to show it the example, I decided to use it here as well.
We can now call our script with the plumb() function, run it with run() and open it on port 800. Calling plumb() creates an environment in which all our functions are evaluated.
library(plumber)r <- plumb("/Users/shiringlander/Documents/Github/shirinsplayground/static/scripts/plumber.R")
r$run(port = 8000)We will now see the following message in our R console:
Starting server to listen on port 8000
Running the swagger UI at http://127.0.0.1:8000/__swagger__/If you go to *http://localhost:8000/__swagger__/*, you could now try out the function by manually choosing values for all the parameters we defined in the script.
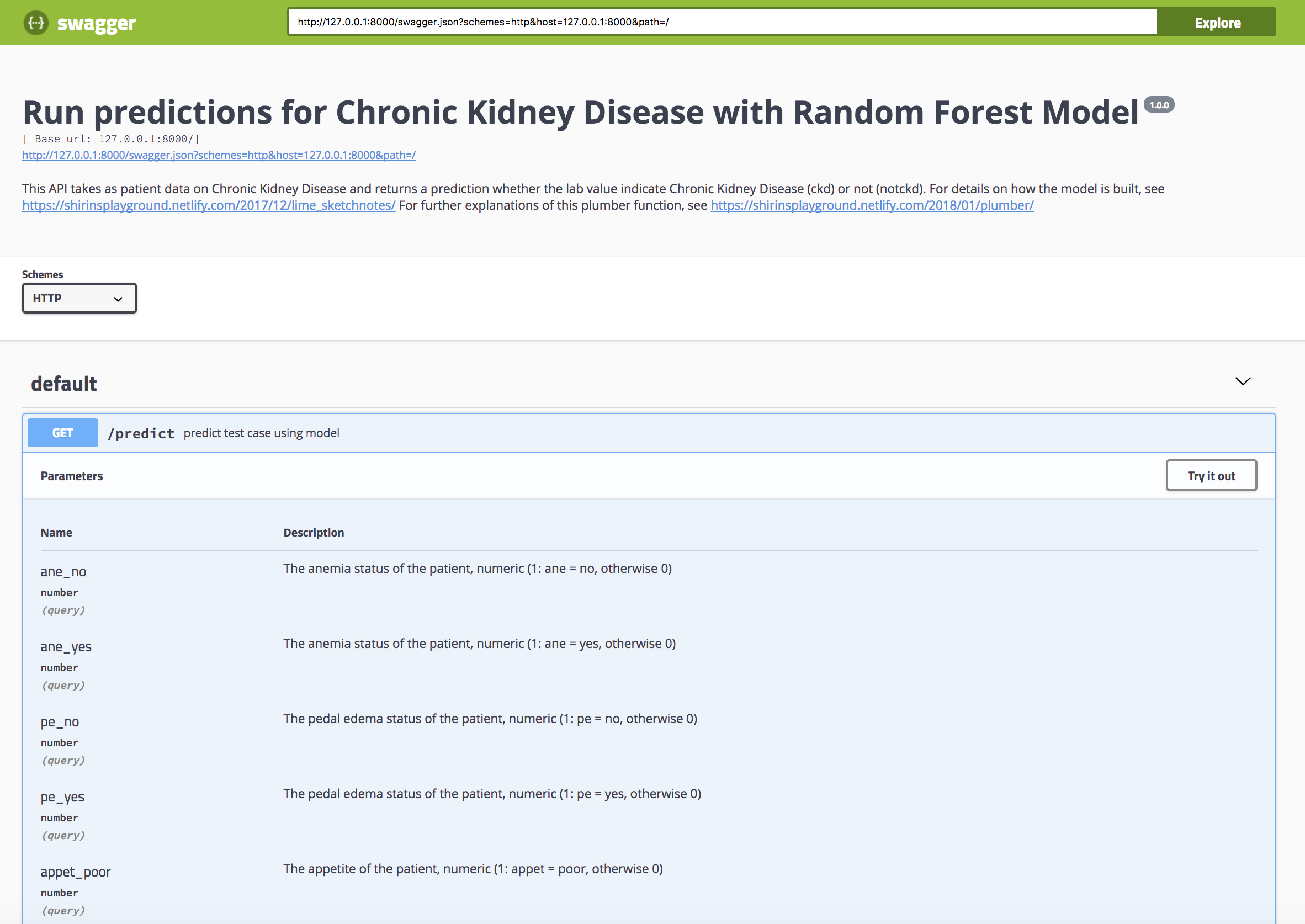 …
… 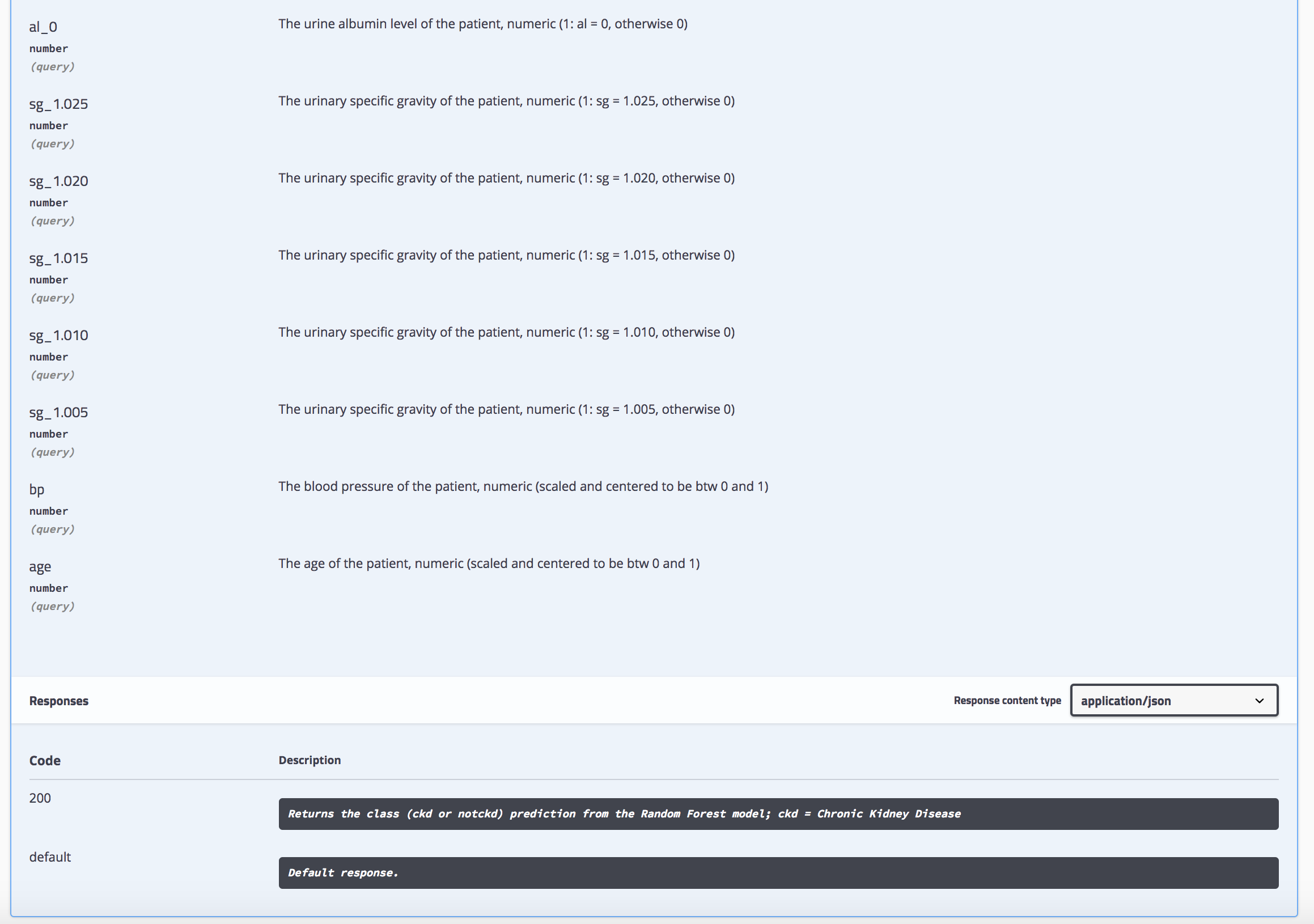
Because we annotated the calculate_prediction() function in our script with #' @get /predict we can access it via *http://localhost:8000/predict*. But because we have no input specified as of yet, we will only see an error on this site. So, we still need to put our JSON formatted input into the function. To do this, we can use curl from the terminal and feed in the JSON string from above. If you are using RStudio in the latest version, you have a handy terminal window open in your working directory. You find it right next to the Console.
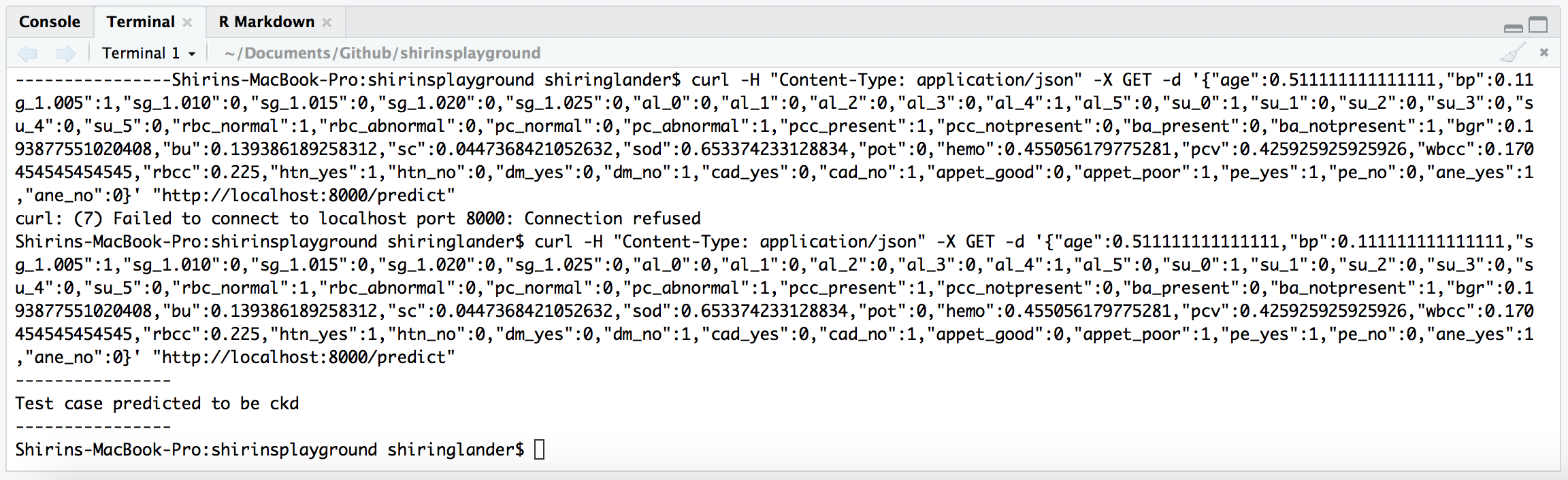
Terminal in RStudio
curl -H "Content-Type: application/json" -X GET -d '{"age":0.511111111111111,"bp":0.111111111111111,"sg_1.005":1,"sg_1.010":0,"sg_1.015":0,"sg_1.020":0,"sg_1.025":0,"al_0":0,"al_1":0,"al_2":0,"al_3":0,"al_4":1,"al_5":0,"su_0":1,"su_1":0,"su_2":0,"su_3":0,"su_4":0,"su_5":0,"rbc_normal":1,"rbc_abnormal":0,"pc_normal":0,"pc_abnormal":1,"pcc_present":1,"pcc_notpresent":0,"ba_present":0,"ba_notpresent":1,"bgr":0.193877551020408,"bu":0.139386189258312,"sc":0.0447368421052632,"sod":0.653374233128834,"pot":0,"hemo":0.455056179775281,"pcv":0.425925925925926,"wbcc":0.170454545454545,"rbcc":0.225,"htn_yes":1,"htn_no":0,"dm_yes":0,"dm_no":1,"cad_yes":0,"cad_no":1,"appet_good":0,"appet_poor":1,"pe_yes":1,"pe_no":0,"ane_yes":1,"ane_no":0}' "http://localhost:8000/predict"-H defines an extra header to include in the request when sending HTTP to a server (https://curl.haxx.se/docs/manpage.html#-H).
-X pecifies a custom request method to use when communicating with the HTTP server (https://curl.haxx.se/docs/manpage.html#-X).
-d sends the specified data in a request to the HTTP server, in the same way that a browser does when a user has filled in an HTML form and presses the submit button. This will cause curl to pass the data to the server using the content-type application/x-www-form-urlencoded (https://curl.haxx.se/docs/manpage.html#-d).
This will return the following output:
cat()outputs to the R console if you use R interactively; if you use R on a server, it will be included in the server logs.
System time: 2018-01-15 13:34:32
Request method: GET /predict
HTTP user agent: curl/7.54.0 @ 127.0.0.1 pasteoutputs to the terminal
----------------
Test case predicted to be ckd
----------------Security
This example shows a pretty simply R-script API. But if you plan on deploying your API to production, you should consider the security section of the plumber documentation. It give additional information about how you can make your code (more) secure.
Finalize
If you wanted to deploy this API you would need to host it, i.e. provide the model and run an R environment with plumber, ideally on a server. A good way to do this, would be to package everything in a Docker container and run this. Docker will ensure that you have a working snapshot of the system settings, R and package versions that won’t change. For more information on dockerizing your API, check out https://hub.docker.com/r/trestletech/plumber/.
sessionInfo()## R version 3.5.1 (2018-07-02)
## Platform: x86_64-apple-darwin15.6.0 (64-bit)
## Running under: macOS High Sierra 10.13.6
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRlapack.dylib
##
## locale:
## [1] de_DE.UTF-8/de_DE.UTF-8/de_DE.UTF-8/C/de_DE.UTF-8/de_DE.UTF-8
##
## attached base packages:
## [1] stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] plumber_0.4.6 rjson_0.2.20 forcats_0.3.0 stringr_1.3.1
## [5] dplyr_0.7.6 purrr_0.2.5 readr_1.1.1 tidyr_0.8.1
## [9] tibble_1.4.2 ggplot2_3.0.0 tidyverse_1.2.1
##
## loaded via a namespace (and not attached):
## [1] nlme_3.1-137 lubridate_1.7.4 dimRed_0.1.0
## [4] httr_1.3.1 rprojroot_1.3-2 tools_3.5.1
## [7] backports_1.1.2 R6_2.2.2 rpart_4.1-13
## [10] lazyeval_0.2.1 colorspace_1.3-2 nnet_7.3-12
## [13] withr_2.1.2 tidyselect_0.2.4 compiler_3.5.1
## [16] cli_1.0.0 rvest_0.3.2 xml2_1.2.0
## [19] bookdown_0.7 scales_0.5.0 sfsmisc_1.1-2
## [22] DEoptimR_1.0-8 robustbase_0.93-2 randomForest_4.6-14
## [25] digest_0.6.15 rmarkdown_1.10 pkgconfig_2.0.1
## [28] htmltools_0.3.6 rlang_0.2.1 readxl_1.1.0
## [31] ddalpha_1.3.4 rstudioapi_0.7 bindr_0.1.1
## [34] jsonlite_1.5 ModelMetrics_1.1.0 magrittr_1.5
## [37] Matrix_1.2-14 Rcpp_0.12.18 munsell_0.5.0
## [40] abind_1.4-5 stringi_1.2.4 yaml_2.2.0
## [43] MASS_7.3-50 plyr_1.8.4 recipes_0.1.3
## [46] grid_3.5.1 promises_1.0.1 pls_2.6-0
## [49] crayon_1.3.4 lattice_0.20-35 haven_1.1.2
## [52] splines_3.5.1 hms_0.4.2 knitr_1.20
## [55] pillar_1.3.0 reshape2_1.4.3 codetools_0.2-15
## [58] stats4_3.5.1 CVST_0.2-2 magic_1.5-8
## [61] glue_1.3.0 evaluate_0.11 blogdown_0.8
## [64] modelr_0.1.2 httpuv_1.4.5 foreach_1.4.4
## [67] cellranger_1.1.0 gtable_0.2.0 kernlab_0.9-26
## [70] assertthat_0.2.0 DRR_0.0.3 xfun_0.3
## [73] gower_0.1.2 prodlim_2018.04.18 broom_0.5.0
## [76] later_0.7.3 class_7.3-14 survival_2.42-6
## [79] geometry_0.3-6 timeDate_3043.102 RcppRoll_0.3.0
## [82] iterators_1.0.10 bindrcpp_0.2.2 lava_1.6.2
## [85] caret_6.0-80 ipred_0.9-6